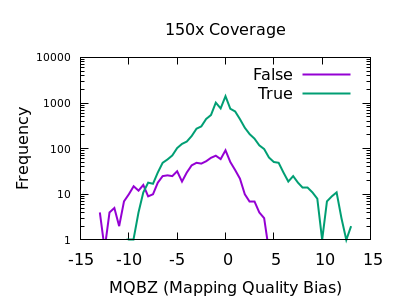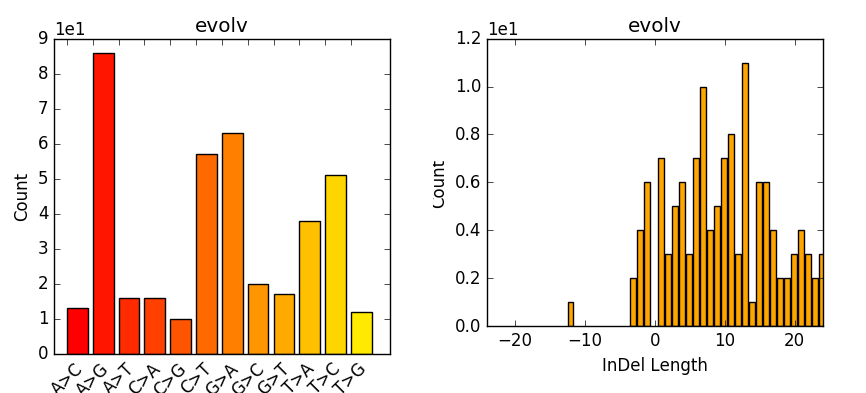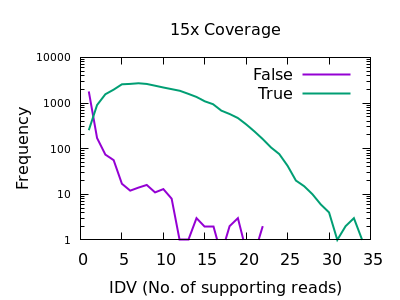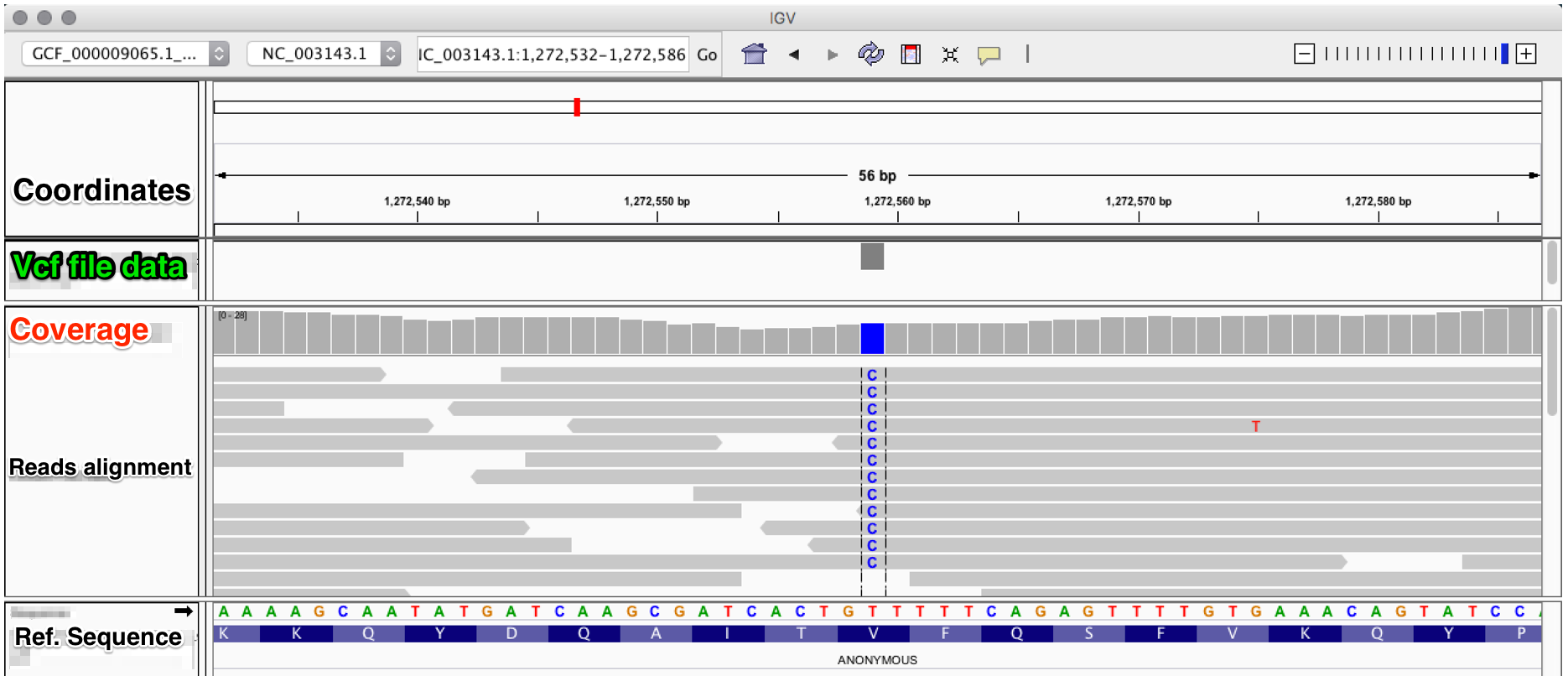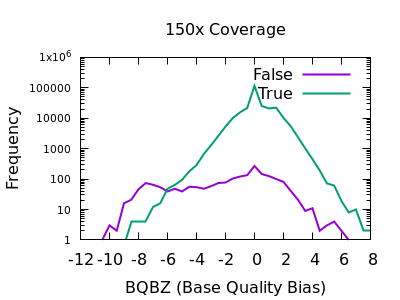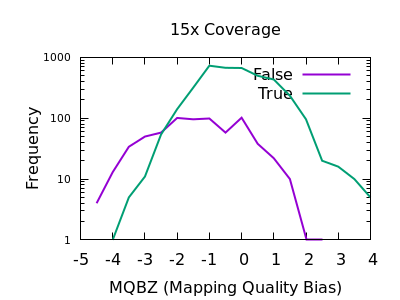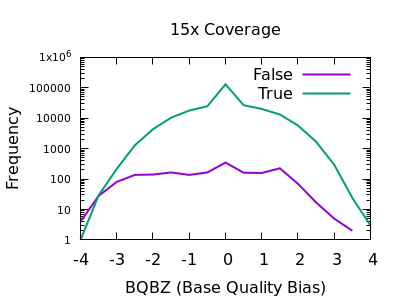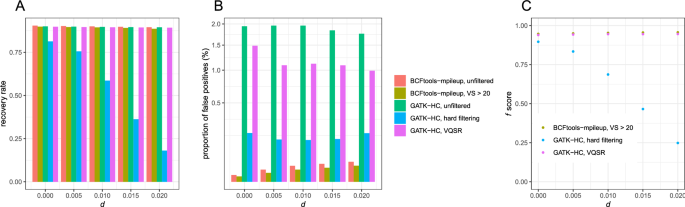
The evaluation of Bcftools mpileup and GATK HaplotypeCaller for variant calling in non-human species | Scientific Reports
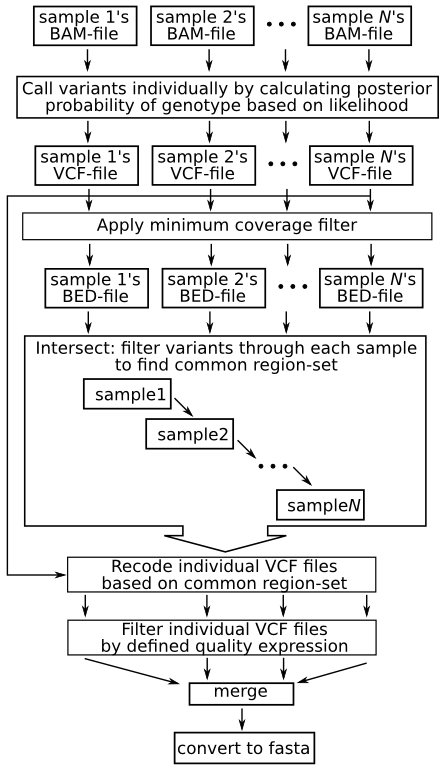
a) Filtering different variant callers VCF output for SARS-CoV-2 data.... | Download Scientific Diagram

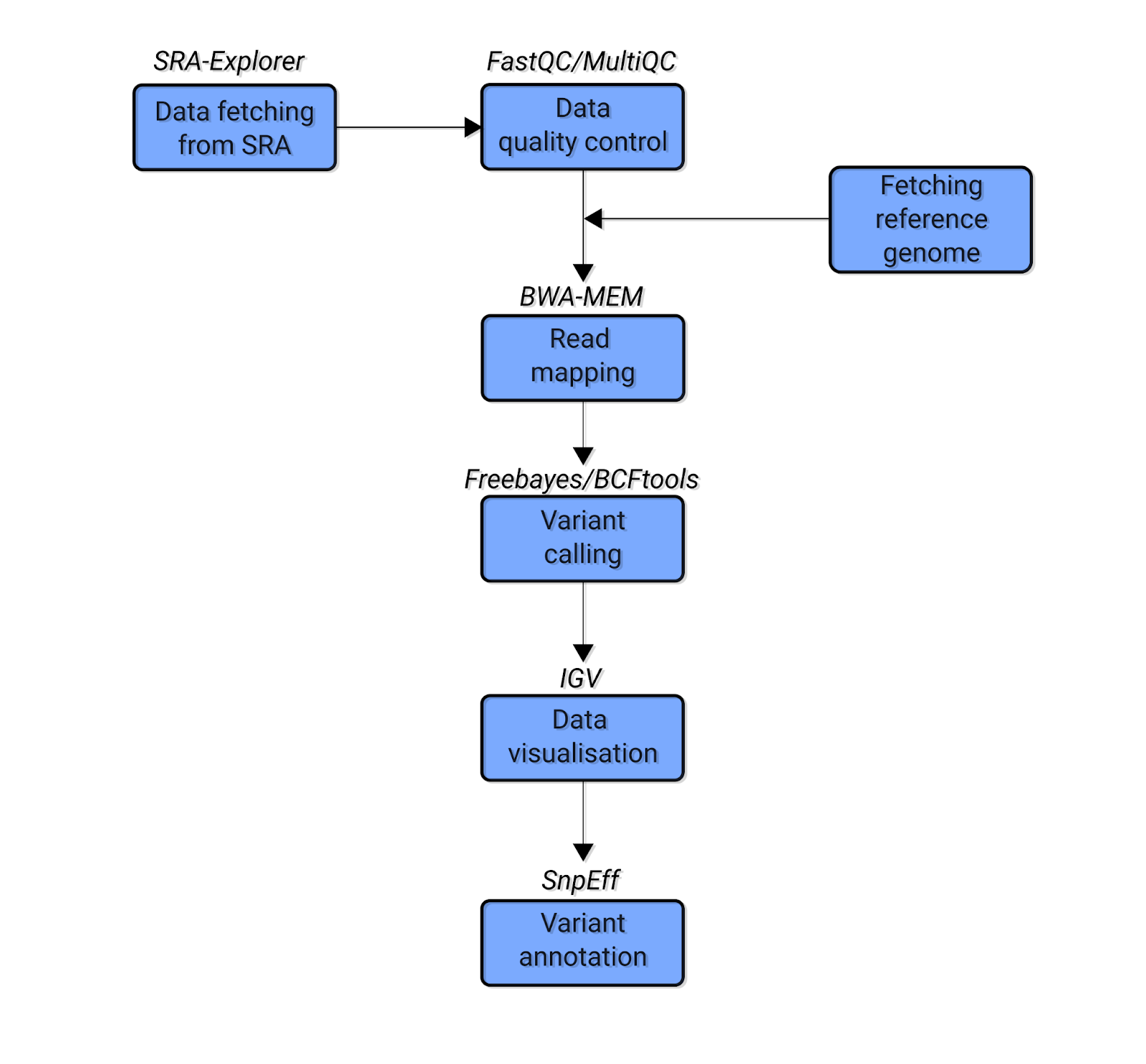
Cells | Free Full-Text | A Next Generation Sequencing-Based Protocol for Screening of Variants of Concern in Autism Spectrum Disorder

The evaluation of Bcftools mpileup and GATK HaplotypeCaller for variant calling in non-human species | Scientific Reports
No unexpected CRISPR-Cas9 off-target activity revealed by trio sequencing of gene-edited mice | PLOS Genetics

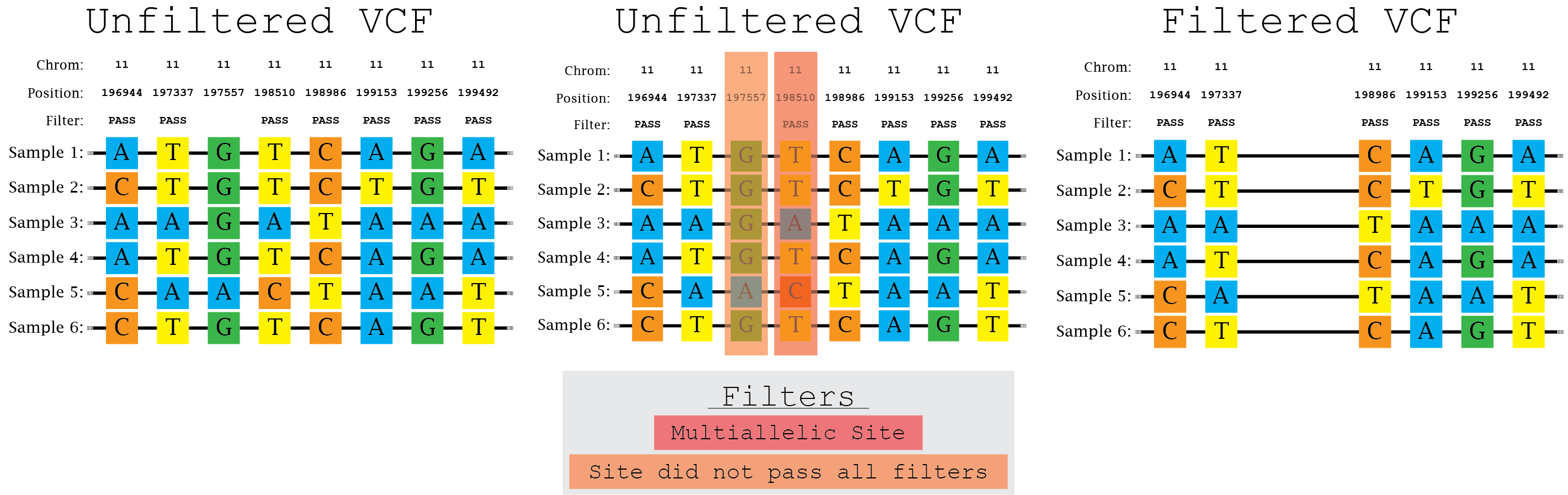
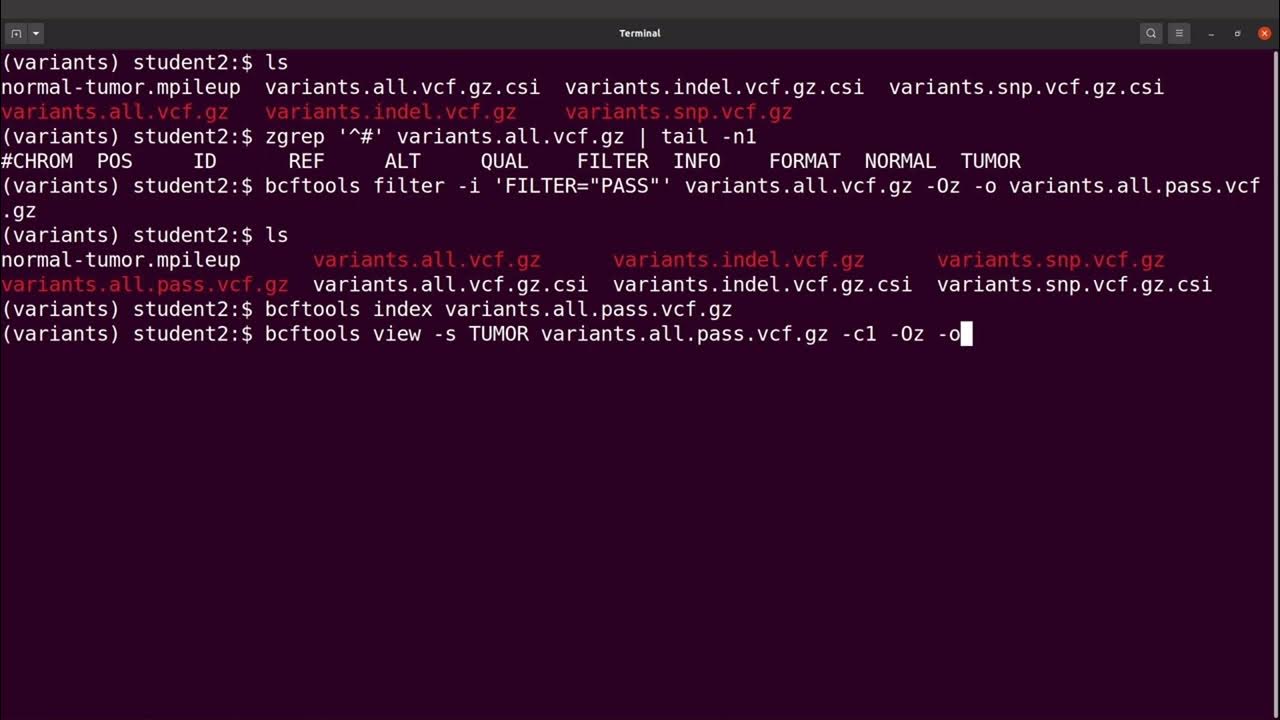
bcftools view | bcftools tutorial on how to count the number of snps and indels in a vcf file - YouTube